ECHO probe
The Best Chemical Probe Spotlighting Nucleic Acid You Want To See
The understanding of the regulation of the mechanisms via which genomic information dictates cellular behaviors has become a great challenge of the postgenomic era. Tools that allow sensitive, quantitative, and real-time detection of specific transcripts, as well as the study of spatiotemporal gene regulation in living cells, are being developed. In this web page, recent advances in nucleic acid detection using exciton-controlled hybridization-sensitive fluorescent oligonucleotide (ECHO) probe technologies are introduced to you. In ECHO probes, a hybridization-dependent fluorescent nucleotide regulated by the H-aggregation of thiazole orange organic dyes (D514) is incorporated into specific sequence contexts and serves as fluorescent detection readout for target nucleic acids. Multicolor detection and auxiliary functional modules have been built into ECHO probes to accommodate a broad range of biological applications.
Principle
ECHO probes have a fluorescence-labeled nucleotide in which two molecules of thiazole orange or its derivatives are linked covalently.
Thiazole orange and its analogues are known to show little emission by the exciton coupling effect when several fluorescent dyes arrange in parallel (H-aggregation). Interaction between the dyes has been explained by Kasha in terms of the exciton coupling theory, in which the excited state of the dye dimer splits into two energy levels. On the basis of exciton theory, the transition to the upper excitonic state is allowed for the H-aggregates, which rapidly deactivate to a lower excitonic state. Emission from the lower energy level is theoretically forbidden and thus the singlet excited state of the aggregate becomes trapped in a nonemissive state.
A bichromophoric aggregate is formed in the ECHO probe and its emission is suppressed strongly by the excitonic interaction between dyes. In contrast, strong emission is observed when the ECHO probe binds to its target nucleic acid, because the aggregate is dissolved and each dye independently intercalates into the nascent duplex structure.
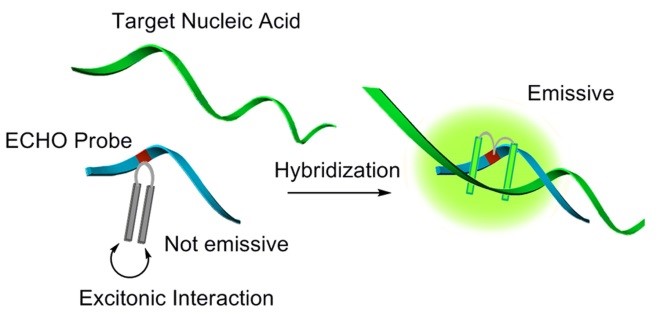
How to choose the sequence
The clear change in fluorescence intensity depending on hybridization is useful for visible gene analysis. However, even this newly designed fluorescent probe exhibits a sequence-dependent change in the ratio of the fluorescence intensities for double- and single-stranded states. The ratio of quantum yields between after and before addition of the complementary nucleic acid strands showed a strong relationship with the number of possible Watson-Crick base pairs formed within two bases from D514 in the ‘self-dimer’ of the probe. The probe self-dimer must be thermodynamically unstable because it contains many mismatched base pairs, but this plausible self-dimeric structure may inhibit fluorescence quenching because the formation of a ‘primer dimer’ in PCR experiments significantly decreases amplification efficiency.
The length of ECHO probes should be 13-30nt. The longer ECHO sequence may form self-dimers and show the strong background fluorescence.
In addition, to choose the sequence where higher-order structure forming or protein-binding region is included, for the ECHO-targeting site, should be avoided. The target of ECHO probes is only the complementary single-stranded nucleic acid. If you have the data on the secondary structure of the target nucleic acid, it would give a hint for the design of probes.
The following sequence design tools provided by DNAForm Co. Ltd. are also very useful for the design of ECHO probes:
http://www.dnaform.com/edesign/
http://www.dnaform.com/echo/thermodynamics/
Technology
Long-term imaging: You can monitor the RNA in a cell for 24-48 h.
Multicolor imaging: You can use the wavelength range of UV to NIR.
SNP analysis: You can detect difference of a single nucleotide.
Highly sensitive analysis: You can see a small amount of expressed RNA.
Aptamer: You can detect the small molecules by ECHO-installed aptamer.
Applications
- Fluorescence in situ hybridization
- Live-cell imaging of mRNA, lncRNA, miRNA…
- Primer for amplification
- Labeling of nucleic acid architectures
- Detection of the molecules recognized by ECHO probes
- SARS-CoV-2 testing kit (SmartAmp assay by DNAForm Co. Ltd.)
… and so on
Availability
D514 ECHO probe is commercially available.
Other colors may also be available. Consult to the suppliers.
- DNAForm Co. Ltd., “Eprobe”, “Eprimer”, or “Exciton primer”.
(https://www.dnaform.jp/en/products/fluorecent_oligonucleotide/eprobe_eprimer/) - Eurofins Genomics Co. Ltd., “Eprobe” or “Eprimer”. Ask sales-jp(at)eurofins.com
- GeneDesign Co. Ltd. (Ajinomoto Bio-Pharma Services), “ECHO probe”. Ask info-e(at)jp.ajibio-pharma.co
m
Papers to be referred
- Chem. Asian J. 2008, 3, 958-968. (First paper)
- Bioconjugate Chem. 2008, 19, 1719-1725. (Sequence choice)
- Bull. Chem. Soc. Jpn. 2009, 82, 110-117. (Live-cell RNA imaging)
- Bioconjugate Chem. 2009, 20, 1256-1261. (Long-term imging by 2’-O-methylRNA backbone)
- Angew. Chem., Int. Ed. 2009, 48, 6480-6484. (Multi-colored ECHO)
- Tetrahedron Lett. 2009, 50, 7191-7195. (Cytosine-typed ECHO)
- Org. Biomol. Chem. 2010, 8, 546-551. (Self-avoidance by ethylcytosine and inosine)
- Hum. Mutat. 2010, 31, 208-217. (Amprification primer for SNP analysis)
- Chem. Rec. 2010, 10, 188-196. (Review)
- PLoS ONE 2010, 5, e13003. (Tag system for sensitive imaging)
- Bioconjugate Chem. 2010, 21, 2276-2281. (LNA use for backbone)
- Org. Biomol. Chem. 2011, 9, 4199-4204. (NIR region)
- Chem. Soc. Rev. 2011, 40, 5815-5828. (Review)
- Org. Biomol. Chem. 2011, 9, 6598-6603. (Intrastrand FRET)
- Bioconjugate Chem. 2011, 22, 1625-1630. (Ratiometric analysis)
- Photomed. Photobiol. 2011, 33, 45-46. (Intrastrand FRET)
- ChemBioChem 2011, 12, 2871-2880. (Caged ECHO and intracellular pinpoint activation)
- Neurosci. Res. 2011, 71, e122. (Fluorescent in situ hybridization)
- RNA 2012, 18, 166-175. (Fluorescent in situ hybridization)
- J. Photochem. Photobiol. C-Photochem. Rev. 2012, 13, 112-123. (Review)
- J. Luminescence 2012, 132, 2566-2571. (Emission mechanism)
- PLoS One 2012, 7, e30236. (Detection of influenza virus by Ishikawa, RIKEN)
- Biochemistry 2012, 51, 6056-6067. (Tool for predicting the thermodynamics by Kimura, RIKEN)
- Org. Biomol. Chem. 2013, 11, 362-371. (pH-sensitive ECHO)
- J. Phys. Chem. A 2013, 117, 27-33. (Fluorescence Correlation Spectroscopy)
- Artif. DNA PNA XNA 2013, 4, 19-27. (Triplex formation)
- Chem. Rec. 2013, 13, 209-217. (Review)
- Methods Mol. Biol. 2013, 1039, 303-318. (Method for caged ECHO)
- PLoS One 2013, 8, e60151. (MGM2 SNP analysis by Shimizu, RIKEN)
- PLoS One 2013, 8, e70942. (Eprobes by Usui and Harbers, RIKEN)
- PLoS One 2013, 8, e73794. (NRF2 SNP analysis by Ishikawa, RIKEN)
- Trends Org. Chem. 2014, 17, 45-57. (Review)
- Methods Mol. Biol. 2015, 1262, 69-87. (Method)
- Neuromethods 2015, 99, 559-584 (Method for ECHO-FISH)
- Bioconjugate Chem. 2015, 26, 412-417. (Compartment-specific imaging)
- Nucleic Acid Res. 2015, 43, e126. (ECHO-liveFISH)
- Oncol. Rep. 2015, 33, 2719-2727. (KRAS mutation analysis by Hanami and Shimizu, RIKEN)
- Bioorg. Med. Chem. Lett. 2016, 26, 4568-4570. (Rapid nuclear import)
- PLoS One 2016, 11, e0146950. (Probe design tool by Kimura and Harbers, RIKEN)
- Chem. Commun. 2017, 53, 9406-9409. (Detection of bacteria ribosome)
- Chem. Lett. 2017, 46, 1803-1806. (Cell uptake)
- PLoS One 2017, 12, e0171225. (HER2 mutation analysis by Usui, RIKEN)
- PLoS One 2018, 13, e0202429. (Detection of fusion genes by Tabe, RIKEN)
- RNA Detection, Humana Press, New York, NY, 2018, 259-272. (Live imaging methods by Ohtan Wang, Kyoto Univ)
- ChemBioChem 2020, 21, 1022-1027. (Live cell sensing of telomerase activity)
- PLoS One 2020, 15, e0243319. (Rapid and ultra-sensitive detection technique by Hanami and Tanabe, RIKEN)

